CMU's COVIDcast Offers Lessons Learned for the Future of Pandemic Forecasting
Media Inquiries
A massive COVID-19 data collection project led by researchers in Carnegie Mellon University's Delphi Research Group(opens in new window) not only helped steer the response to the virus but could also change the future of pandemic forecasting, the researchers involved concluded.
In a collection of articles published in the Proceedings of the National Academy of Sciences, the researchers behind the COVIDcast repository, the US COVID-19 Trends and Impact Survey (CTIS), and the international version of CTIS reflect on and share lessons learned from their first year collecting data about the pandemic. PNAS featured the project on its January 2022 cover and included the research in a collection, "Beyond Cases and Deaths: The Benefits of Auxiliary Data Streams in Tracking the COVID-19 Pandemic(opens in new window)."
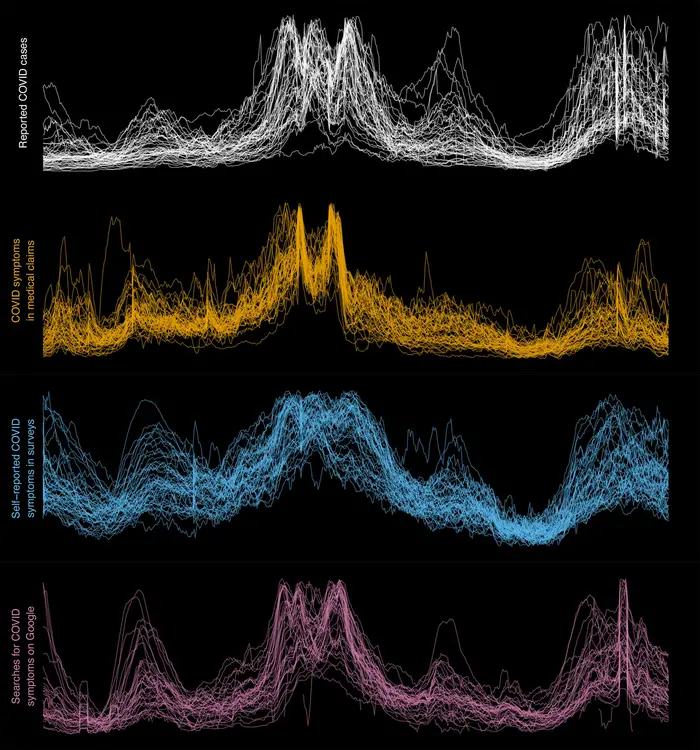
Pictured are plots (from top to bottom) of reported COVID-19 cases, the fraction of doctors' visits with diagnostic codes indicating a COVID-like illness, the fraction of survey respondents who knew somebody with a COVID-like illness, and a measure of internet search volume for specific disease-related terms for each U.S. state from April 2020 to October 2021, scaled to the same dynamic range. Image courtesy of Ryan J. Tibshirani
"When the pandemic broke out, we wanted to find a way to contribute to national efforts to respond," said Roni Rosenfeld(opens in new window), head of the Machine Learning Department(opens in new window) at CMU and a co-director of the Delphi Research Group. "We put our focus on the data, building and making publicly available tools and new indicators that reflected the pandemic in real-time, aided in urgent decision-making and shaped the way public health data could be used in the future."
Through the COVIDcast repository, the researchers compiled a diverse set of real-time, geographically detailed data different from that collected by typical public health reporting. The repository is unique in its breadth, depth, scope and timeliness. The effort attempts to fill the needs of policy makers, epidemiological modelers and health researchers who require up-to-date data on the pandemic and public behavior related to it.
One of the collection's papers, "An Open Repository of Real-time COVID-19 Indicators(opens in new window)," details the COVIDcast repository's impact. COVIDcast receives data daily from health care systems, tech companies, testing results, insurance claims and surveys. Much of this data is unique to COVIDcast, and is produced through collaborations with companies such as Change Healthcare, Facebook and Google. The repository handles hundreds of thousands of requests from thousands of users a day, helping with decision-making, forecasting, and studying the impacts of COVID and the effectiveness of interventions aimed at stopping it.
"Collecting and combining this data is no small feat on its own," said Alex Reinhart(opens in new window), an assistant teaching professor of statistics and data science(opens in new window) at CMU, member of the Delphi Group, and lead author on the paper. "But providing convenient, real-time access to this data enables us to see how things are, how they are expected to change, where we should allocate resources and the effectiveness of communication."
The role of indicators outside typical public health data is further explored in another paper, "Can Auxiliary Indicators Improve COVID-19 Forecasting and Hotspot Prediction?(opens in new window)" The study examined five indicators gathered from deidentified medical insurance claims, self-reported symptoms from online surveys, and Google searches for information about the loss of taste and smell, and found that each indicator increased the accuracy of traditional pandemic forecast models.
The collection also includes a paper about the US COVID-19 Trends and Impact Survey (CTIS)(opens in new window), in which researchers discuss how more than 20 million responses from more than 350,000 people every week informed public health actions. The University of Maryland runs an international version of the survey, which is discussed in a separate paper(opens in new window). These projects are the largest domestic and international public health surveys to date.
Throughout the PNAS collection, the researchers reflect on their first year of data collection and analysis and share lessons they learned. Researchers found that good data on vaccine acceptance can inform policies aiming to increase vaccine uptake, and suggest the data will be useful to future vaccination campaigns. Researchers also expect similar online surveys to play increasingly important roles in future epidemics and pandemics by supplementing public reporting systems with information that is difficult to gather any other way.
In "Epidemic Tracking and Forecasting: Lessons Learned From a Tumultuous Year(opens in new window)," Rosenfeld and Ryan J. Tibshirani(opens in new window), a professor of statistics and machine learning at CMU and co-director of the Delphi Group, reflect on the project as a whole. The pair discuss the importance of specific, clear and consistent data labels, pointing out that terms like COVID cases, hospitalizations and deaths hide enormous amounts of complexity and potential ambiguity. They also highlight the difficulty of measuring and modeling human behavior despite the impact it had on the progression of the pandemic. Behavior like compliance with policies and recommendations are not regularly captured in publicly available data. But that is only half of the challenge.
"Even with this data, we will need significant and new cognitive and behavioral modeling to use it successfully," Tibshirani said. "And how do you gather data on the impact of the breakdown and fragmentation of trust in governments, public health officials and health care professionals? This is difficult to measure and model, but affected the pandemic worldwide."
The full collection of papers is available on the PNAS website(opens in new window). Information about COVIDcast, CTIS and links to the respective dashboards are available on the Delphi Group's COVID-19 page(opens in new window).