
Strength in Numbers
by Amy Pavlak
For more than one hundred years scientists have studied microorganisms as individual cells floating in solution. But microbes are much more likely to exist in communities that adhere to surfaces, from rocks in a stream and the enamel of our teeth to medical implants like heart valves and artificial joints. Scientists at Carnegie Mellon are unraveling the complex structure and biology of these microbial cities and are developing new strategies to combat those that are dangerous.

From left: Fred Lanni, Luisa Hiller, Aaron Mitchell and Alan Russell
Fred Lanni carefully removes the postage-stamp-sized piece of silicone from its watery home in the microwell plate. He beckons me to look closely, and I can just barely see what appears to be white fuzz growing on the silicone’s surface. It looks innocuous enough, but Lanni assures me that there’s much more to the fuzz than the naked eye can see. Images of the fuzz taken with a confocal microscope reveal a thriving microbial city teeming with fungal cells that are layers deep. I was looking at a biofilm, a fortified community of microbes that can grow on nearly any surface—rocks in a stream, a ship’s hull, your teeth, a child’s middle ear, an artificial hip, or a silicone catheter.
For much of the past century, scientists have studied microbes—single-celled organisms like fungi and bacteria—as free-floating cells that they isolated, cultured and suspended in a liquid. But over the past few decades, microbiologists have begun to appreciate that the majority of microbes live in biofilms. And the need to understand these microbial communities is urgent. Biofilm-based microbes are responsible for a wide range of chronic, difficult-to-treat infections.
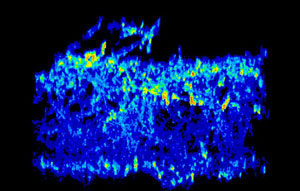
Gene expression pattern of two genes (red/yellow and light blue) active in a C. albicans biofilm. By locating where in the biofilm certain genes are expressed, Mitchell and colleagues are gaining insight into the genes’ function. Images courtesy Fred Lanni and Aaron Mitchell.
Carnegie Mellon Biological Sciences Professor Aaron Mitchell has been studying biofilms for the past 15 years. Four years ago he teamed up with Lanni, an associate professor of biological sciences and an expert in imaging technologies. Their collaboration led to the development of techniques that allow the scientists to image deep inside biofilms, particularly those formed by the fungus Candida albicans. C. albicans biofilms are commonly found on the surfaces of implanted medical devices such as venous catheters, dentures, and urinary catheters. The biofilms cause stubborn infections that can become nearly impossible to treat with common antimicrobial drugs.
“Microbes living in a biofilm are covered in this slimy stuff called biofilm matrix material, and it has profound biological impact. It can prevent drugs from reaching the microbial cells living deep inside the biofilm,” Mitchell said.
Many thousands of microbes can live in this slimy matrix, which provides structure, nutrients and a venue for the cells to communicate with one another.
“Biofilms are as complex as an entire city, and their performance is incredible—they can change their characteristics in order to retain the biofilm in the face of a changing environment. It’s just phenomenal,” said Alan Russell, the Highmark Distinguished Career Professor and director of Carnegie Mellon’s Disruptive Health Technology Institute. Russell’s work focuses on designing ways to prevent or disrupt bacterial biofilm growth on certain surfaces such as catheters and teeth. He’ll be the first to tell you that it has been a vexing challenge.
“Bacteria have developed biofilms for a reason,” Russell said. “They want to survive and thrive, and they’ve had a few billion years to figure out how to do it.”
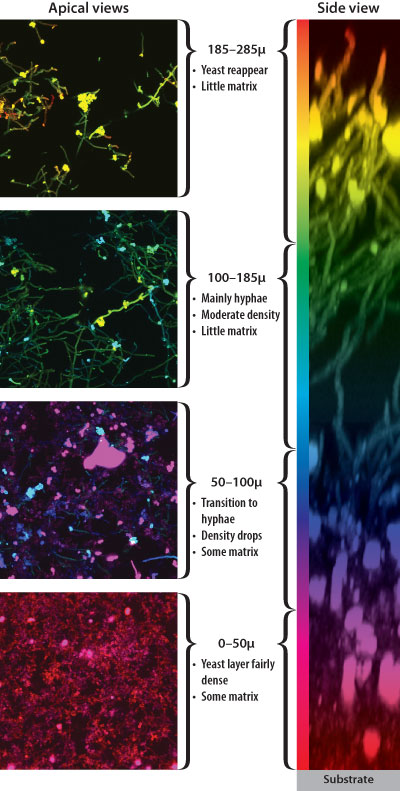
Biofilms are made up of many layers of microbes, the composition of which can vary from the top to the bottom. Images courtesy Fred Lanni and Aaron Mitchell.
Biofilm formation begins when free-floating microbes, such as those in saliva or the air, settle on a surface, grow into a cluster and then stick. The cluster of microbial cells begins producing the gooey extracellular matrix material, providing a protected environment that the microbes control.
To understand the mechanisms by which C. albicans biofilms develop and grow, Mitchell and his colleagues compare the gene expression patterns of fungal cells growing in a biofilm to those that are suspended in liquid. It turns out that fungi’s living arrangements play a big role in what genes they express.
Working in collaboration with alumnus Vincent Bruno (S’96), an assistant professor at the University of Maryland School of Medicine, the researchers have identified several genes that are more highly expressed in the biofilm cells, including some that are involved in biofilm formation. In their most recent work, the Mitchell lab discovered that a gene called RHR2 was one of the most highly expressed in fungal biofilm cells relative to free-floating cells. The RHR2 gene encodes an enzyme that synthesizes glycerol, a small metabolite that is abundant in biofilm cells. When graduate student Jigar Desai (S’14), who works with Lanni and Mitchell, started looking at RHR2 more closely, he found that glycerol is required for the expression of numerous biofilm-associated genes, including genes responsible for cell surface adhesion, a key step in biofilm formation.
To test RHR2’s role in adhesion, Desai engineered fungal cells that don’t have the RHR2 gene. He then tested the mutant cells’ ability to grow on silicone—one of the primary surfaces to which C. albicans cells stick, and the main material in many medical implants. Desai found that the fungi without RHR2 didn’t adhere as well to silicone and formed less biofilm than their fully functional counterparts. The promising results prompted Mitchell to send the mutant strain of RHR2-deficient fungal cells to David Andes, M.D., a professor in the Departments of Medicine and Medical Microbiology and Immunology at the University of Wisconsin. Andes tested the mutant cells in an animal model of catheter infection. He found that the fungal cells lacking RHR2 almost completely lose their ability to form a biofilm. As Mitchell and his team continue to uncover more about RHR2, their excitement builds.
“It suggests that metabolic enzymes may have broad regulatory effects on biofilm formation, and that they might be excellent targets for therapeutic development,” Mitchell said.
The success of microorganisms living in biofilms depends on their ability to communicate with one another. In bacterial biofilms, the bacteria send chemical signals that allow them to sense each other’s presence and respond. These conversations allow the bacteria to take on different tasks from ferrying nutrients and removing wastes to taking up DNA from the biofilm’s slimy matrix and using it to modify themselves.
“One thing that is noteworthy about biofilms is that they all have extracellular material. The exact material varies, but there’s one constituent that is found in almost every biofilm—extracellular DNA,” Mitchell said.
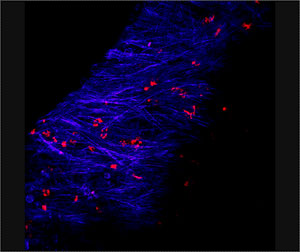
Micrograph of a chronic biofilm infection caused by Streptococcus pneumoniae in a chinchilla ear. Blue denotes chinchilla tissue and red denotes bacteria. Image courtesy Laura Nistico.
The bacteria that commonly cause middle ear infections in children, Streptococcus pneumoniae (pneumococcus for short), like to form biofilms. When in a biofilm mode of growth, these bacteria are up to 1,000 times more resistant to antibiotics. Antibiotic resistance is one of the reasons persistent childhood ear infections are so difficult to treat. Treatment options are further complicated by pneumococcus’s skill at taking up extracellular DNA and incorporating it into its own chromosome. This capability allows the bacteria to easily acquire new traits from their neighbors in the biofilm—such as antibiotic resistance and the ability to engage in new interactions with their human host.
Luisa Hiller, assistant professor of biological sciences, studies pneumococcus’s remarkable ability to take up extracellular DNA. As a postdoc, she looked at “pneumo” strains isolated from one child with a chronic, biofilm-based ear infection. She found that, over seven months of infection, one of the strains infecting the child acquired about 10 percent of its genome from another strain in the same biofilm. In genome terms, that’s equivalent to a mouse becoming a human. The genes the strain picked up provided different features, such as a new bacterial coat.
In recent work in her lab, Hiller has begun digging into the mechanisms that control the extent to which these bacteria take up DNA. Hiller is comparing the entire genomes of different strains of pneumococcus, particularly those that tend to take up a lot of DNA and those that don’t. Computational analyses are revealing genomic differences across strains, which can help to zero in on which genes are present in bacteria that are adept or inept at taking in DNA.
Hiller is also interested in how the individual bacterial cells in a biofilm talk to each other. Sometimes the conversations are friendly; other times they are belligerent. Anagha Kadam, a graduate student in Hiller’s lab, studies molecules that carry on such conversations. Working with bacterial samples from the ear of a chinchilla, an animal model of ear infection, Kadam looks at the RNA levels inside the bacterial cells. She found that certain molecules, which Hiller calls war molecules, are highly expressed during the infection.
“Pneumo has a lot of molecules that seem to be devoted to warfare, including warfare against slightly different strains of their own species,” Hiller explained. She points out that many strains of pneumococcus are present at the same time in a biofilm, so the “hypothesis in the field is that, in times of stress, you kill some neighbors and decrease competition for resources. But you also get their DNA. And that increases the probability of now reaching a new genome that’s better adapted.”
As Mitchell and Hiller work to unravel some of the complex behaviors microbes exhibit while living in a biofilm, they also are thinking about ways to kill them—or at least prevent their formation.
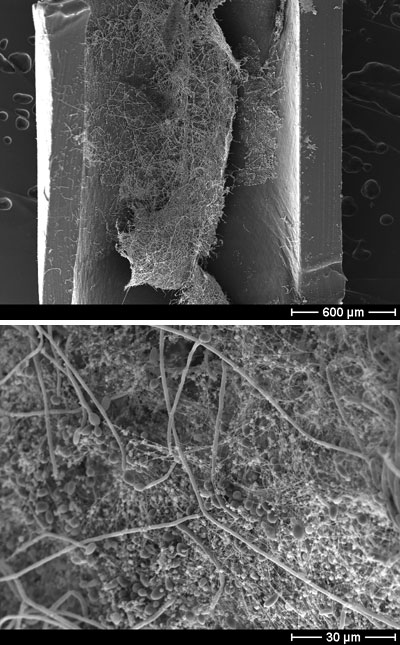

Scanning electron microscope image of a C. albicans biofilm growing in an intravenous catheter. (Left) 50x magnification. (Right) 1000x magnification, which reveals the biofilm’s constituent parts—oval-shaped yeast cells, filamentous hyphae, and stringy extracellular matrix. These experiments were carried out by Mitchell’s collaborators, Kaitlin Mitchell and David Andes, at the university of Wisconsin. Image courtesy Aaron Mitchell.
“We’re definitely interested in how biofilms form and behave. That’s kind of the basic science angle,” Mitchell said. “We’re also working on a project with Fred [Lanni] that is much more applied. In that case we actually don’t care how biofilms form or behave. We just care that we can kill them in a broadly useful way.”
Hiller, Mitchell and Lanni are taking a cue from a type of cancer treatment known as photodynamic therapy to develop a recyclable anti-biofilm coating for medical devices. This method for treating tumors uses certain dyes that, when hit with light, generate a toxic form of oxygen called singlet oxygen that kills tumor cells. Lanni reasoned that the same dyes could be used to kill microbes that adhere to surfaces before they have a chance to form a persistent biofilm. The three are developing a coating containing a nonreleasable dye, which ultimately could be applied to the surfaces of medical devices like venous catheters, bone pins in extremities or possibly even deeper hardware such as joint implants. Treating the patient externally and locally with deep red light, which penetrates tissues well, would cause the dyes to generate singlet oxygen, killing any microorganisms that had adhered to the medical device. In principle, this could be done chronically to suppress infection during the treatment or healing period.
The research team has performed in vitro tests to evaluate the dyes’ ability to kill fungi and bacteria, which have proven successful. The next step is to do testing in an animal model.
The method is particularly appealing for a few reasons. Singlet oxygen species have a very short half-life, so they can’t diffuse very far away from the dye. This keeps their killing power localized. Additionally, one dye molecule can activate many oxygen molecules and can be hit with light again and again, creating a way to keep the medical implant free of biofilms potentially for the life of the implant.
“You don’t want to take out your artificial hip every year or so for a new treatment. You want a lifetime warranty,” Mitchell said.
Russell, who works at the interface of chemistry, biology and materials, has taken a different approach to anti-biofilm coatings. He recently developed an aspirin-releasing polymer that coats the inner lumen of urinary catheters. Inspired by the fact that aspirin has the ability to disrupt biofilms, Russell and his team made a polymer that releases aspirin over time. The catheter, which the pharmaceutical company Bayer has patented, reduces biofilm formation by E. coli for up to five days under conditions that simulate urine flow.
Russell’s group is currently working on a project to coat medical implants with polymers paired with probiotic (“good”) bacteria. “The question is: can you create a biofilm that’s good? Can we design surfaces that will attract good biofilms versus bad ones?” Russell said.
He maintains that biofilms are a career-sustaining area.
“This is a field where you take three steps forward and two steps back and you keep doing that and you advance yourself. But each time you take a step back, you have to learn why. You have to understand biofilms if you’re going to prevent their growth. And that’s why CMU has something really interesting, I think. We have a really hard-core depth in the science of biofilms and we have a lot of people on campus who understand how to put biology and chemistry and materials together.”