A World of Differences
Biologist Veronica Hinman scans entire genomes to pinpoint the molecular processes that have generated the great diversity of animal forms found in the world today.
by Amy Pavlak
Veronica Hinman did her first dive in the Great Barrier Reef when she was a teenager, strapping on her scuba gear and descending into an underwater paradise teeming with life. Like many divers exploring the 1,400-mile-long reef, she was astounded by the extraordinary diversity of creatures living in the clear, warm waters.
The budding scientist in her wondered how the tens of thousands of creatures living in the reef got to be so different—and she’s devoted her career to finding out. In the almost thirty years since Hinman made her first dive in the Great Barrier Reef, she has been on a quest to answer one of the most intriguing questions in biology: How can so much diversity be produced from the same basic genetic toolkit that’s shared by all animals? For Hinman, an associate professor of biological sciences, the answer may lie in studying some of the very same sea creatures that inspired her as a teenager.
Growing up in a fishing village in Portugal and then in Brisbane, Australia, Hinman has always had an affinity for the sea. She also had an inquisitive mind and an interest in science, especially evolution. As an undergraduate at the University of Queensland, she took classes in paleontology and engineering, but neither field struck her fancy. It wasn’t until her junior year, when she took an ecology and evolution class, that she found her niche. And it didn’t hurt that she was at one of academia’s largest marine labs.
“The marine environment is where it’s at,” Hinman says. “At marine stations, there’s always people working on a great diversity of organisms. It was so exciting to see people who were really into the biology, the diversity. It was a really interesting place.”
Hinman stayed at the University of Queensland to pursue a Ph.D. in zoology. She cut her teeth as a young graduate student during a revolution in evolutionary biology. It was the mid-1990s and scientists had recently discovered something surprising—humans and mice and fruit flies had the same genes. Given the diversity of organisms on land and in the sea, scientists expected their genes to be just as diverse. The new findings blew everyone’s minds, including Hinman’s.
“There’s no way that anyone was thinking that a fruit fly and a human would have the same genes. We’d never, ever have guessed that this would be the case.”
Hinman began thinking in earnest about the question she first pondered while diving the Great Barrier Reef: How could these animals be so different if they share the same genes? She decided to start at the beginning and try to understand how animals take on their form during embryonic development.
Every animal starts as an egg, goes through embryogenesis and becomes an adult. So if you’re going to change what they become, explains Hinman, you have to change how they develop. She realized that to truly understand the mechanisms that control development, she needed to become rigorously trained in molecular biology and biochemistry. She sought out a postdoctoral position at the California Institute of Technology with Eric Davidson, who ran a lab focusing on molecular genetics. Hinman delved into how genes were being regulated during the early stages of sea urchin embryo development.
“I didn’t necessarily mean to keep the marine theme. It just sort of happened that way,” Hinman says. “But I think the reason is because the sea urchin is a very simple organism, so you can really understand very precisely how development proceeds.”
As embryos cleave from one cell into two, and then two into four, multiple genes are switched on and off in a very precise way to form every aspect of the embryo and eventually its adult form. Despite their ultimate differences in appearance, all complex animals have the same set of homologous genes called the “developmental gene toolkit” that carry out this finely tuned dance.
“When we’re trying to understand evolution, it’s not so much the genes themselves, then, but how they’re regulated,” Hinman said.
Since joining the Carnegie Mellon faculty in 2006, Hinman has been exploring precisely how genes are being regulated in developing embryos. The advent of fast, inexpensive sequencing technologies is allowing Hinman to probe the genomes of sea urchins and their relatives in ways that weren’t possible just a few years ago. Using a relatively new technique called chromatin immunoprecipitation with deep sequencing, she can quickly sequence DNA from thousands of embryos and then scan the entire genome to find regions that regulate how specific genes are deployed: when and where they are turned on, for how long, and to what levels.
“Now you can look everywhere—all those regions of the genome that maybe you weren’t even looking at before.”
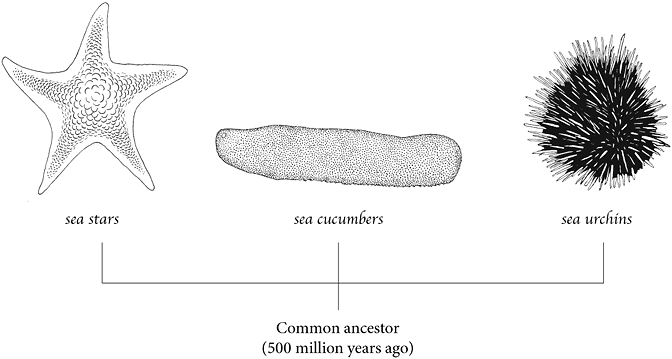
Hinman is comparing the genomes of sea urchins, sea stars and sea cucumbers, which are all descended from an ancient lineage going back 500 million years. Although they share a common ancestor, each went down different evolutionary paths to become distinctly different animals. Hinman is teasing apart their entire genomes to determine how their DNA evolved to produce such astoundingly different creatures.
Hinman is particularly interested in understanding a marked change that occurs in an early stage of embryo development—sea urchin larvae form complex skeletons, sea cucumbers form teeny, tiny skeletons and sea stars don’t form skeletons at all. Hinman is comparing their genomes to pinpoint how the regulatory regions of the DNA have evolved to allow such differences to happen.
For Hinman, it’s come full circle from her early days as a graduate student when she and her colleagues were fascinated that animals shared the same genes. Now, the focus has turned to understanding what makes animals different.
“I believe that during my lifetime we will be in a position to really understand how animals are made and most importantly how they are made in similar and different ways.”
