
Biological Physics
Supramolecular Structures Lab
Lösche/Heinrich Group
Research
Sparsely-tethered Bilayer Lipid Membranes (stBLMs) for Biomedical Research
Physiological bilayer membranes – fluid leaflets self-assembled by non-covalent interactions and a mere 5 nanometers thin – are ubiquitously used in biological cells but much to delicate to use in practical applications. While a great deal of fundamental research has been achieved with model systems such as "Black Lipid Membranes", i.e. freely suspended bilayers between two semi-infinite buffer compartments, these are entirely unsuitable if it comes to long-term experiments, be it systematic screening, observations over extended periods or technological applications. Also, such BLMs are not suitable for high-res structural investigations, because they're so small in lateral extension (~ 10-2 mm2).
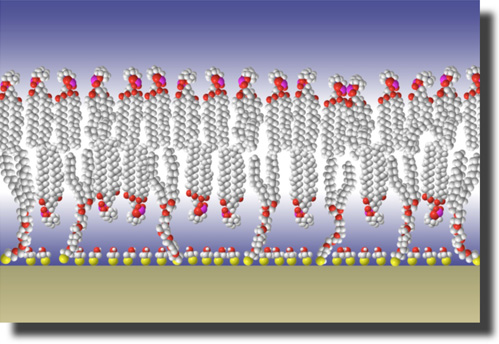 Self-assembled lipid membranes may serve as functional interface architectures in applications that range from biosensorics to pharmacological screening.
Self-assembled lipid membranes may serve as functional interface architectures in applications that range from biosensorics to pharmacological screening.
There are established techniques to form, or to transfer, bilayer membranes on solid supports. These approaches all have the potential to address both the stability and the size limitation issues. However, membranes in contact with solid substrates loose their intrinsic dynamics – both in-plane and, of course, out-of-plane – due to the interaction of the constituent lipids with the solid surface. Also, proteins which one might want to reconstitute into lipid membrane for functionalization, tend to denature (unfold from their specific three-dimensional structure required for protein function) at the surface.
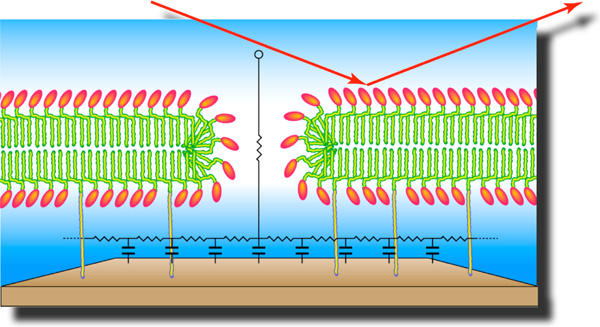 Our workhorses: Neutron or x-ray reflection is used to characterize the molecular structure of stBLMs. Electrochemical characterization is used to assess their capacitance, residual conductance, and defect density. Fluorescence correlation spectroscopy (FCS) is a useful technique to study membrane dynamics and surface plasmon spectroscopy (SPR) quantifies the kinetics of protein or peptide binding to functionalized membranes.
Our workhorses: Neutron or x-ray reflection is used to characterize the molecular structure of stBLMs. Electrochemical characterization is used to assess their capacitance, residual conductance, and defect density. Fluorescence correlation spectroscopy (FCS) is a useful technique to study membrane dynamics and surface plasmon spectroscopy (SPR) quantifies the kinetics of protein or peptide binding to functionalized membranes.
We have developed specific molecular surface architectures, substrate-supported tethered bilayer lipid membranes (tBLMs), in which the 50 Å thick bilayer membrane is adsorbed to a solid surface via a hydrated chemical tether. Ingredients for this work are a dedicated synthetic chemistry (developed by David Vanderah at NIST-CSTL), an unconventional preparation protocol for bilayer completion (first introduced by the Cornell group), electrochemical impedance spectroscopy (the EIS technique has been brought to us by Gintaras Valincius of the Biochemistry Institute in Vilnius, Lithuania) for facile system optimization, and neutron reflectometry (NR) for the high-res structural assessment of the resulting architecture.
The result is an optimized experimental system, easy and reproducibly to prepare in simple bake-and-shake sample preps, that we use in various applications within a multitude of collaborations:
– The pathologically large association of Lamin A mutants with models of the nuclear membrane (with Kris Dahl at CMU Biomedical Engineering).
– Structural details of protein-membrane association in the HIV infection pathway (with Alan Rein at the NIH-NCI and Volker Vogt at Cornell).
– Cellular control of the tumor suppressor PTEN (with Arne Gericke at Worcester Polytech and Alonzo Ross at the UMass Med School in Worcester, MA).
– Association of neurotransmitters with lipid membranes (with Robert Cantor, Dartmouth)
– Characterization of the cellular entry mechanism of the endolysin, PlyC (with Dan Nelson, U Maryland/IBBR)
– Endosomal release of diphtheria toxin (with Alex Ladokhin, U Kansas Med School)
In all cases, the tBLMs permit us to explore dimensions in biomedical research that have not previously been accessible.
Research Wiki
A collaborative platform for research
(internal access only)
Research topics
stBLM optimization at the NIST Center for Neutron Research. More...
Alzheimer's peptide amyloid β: Mechanisms of Aβ toxicity. More...
Initial steps of viral assembly studied in structural investigations of the HIV-1 gag protein at membrane surfaces. More ...
Membrane incorporation of the protein toxin pore α-hemolysin. More...