Biological Physics
Supramolecular Structures Lab
Lösche/Heinrich Group
Supramolecular Structures Lab
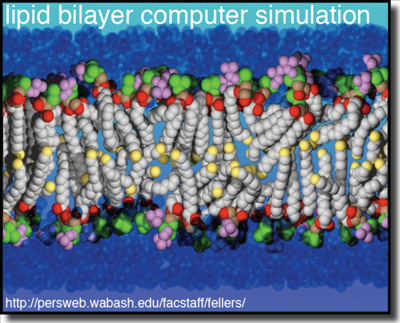
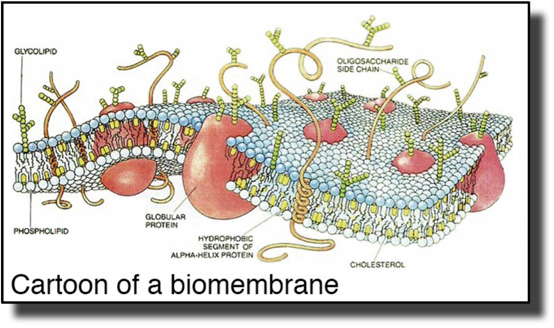 Ordered molecular assemblies form the structural basis of life. Their length scales range from nanometers, the thickness of a biomembrane, to micrometers, the length of polymers that form the cytoskeleton. They are chemically diverse – building blocks include proteins, nucleic acids, lipids and polysaccharides – and formidably complex.
Ordered molecular assemblies form the structural basis of life. Their length scales range from nanometers, the thickness of a biomembrane, to micrometers, the length of polymers that form the cytoskeleton. They are chemically diverse – building blocks include proteins, nucleic acids, lipids and polysaccharides – and formidably complex.
As are the underlying interactions: Van-der-Waals, electrostatic, hydrogen bonding and hydrophobic forces all contribute to their specific organization. Since these interactions are characterized by thermal energies, the resulting structures are highly dynamic, capable of reorganizing in response to a changing environment.
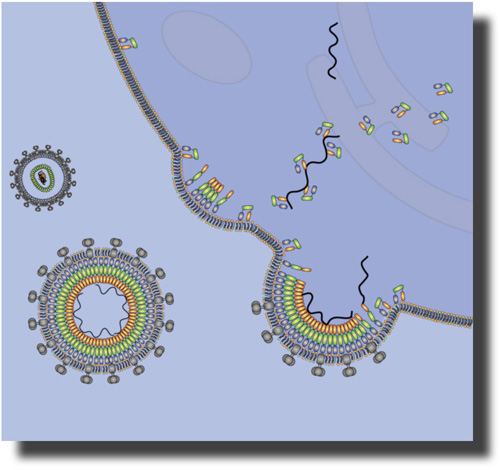 We are particularly interested in biomembranes and membrane-related biological processes. For example, many toxins exert their damage on target cells by interfering with membrane functions. Misfolded peptides affect membranes of neuronal cells so that the communication between these cells is abolished; diseases such as Alzheimer's or Parkinson's are the consequence. As a further example, viral assembly, such as that of the Human Immunodeficiency Virus occurs at the plasma membrane of host cells and is a complex process involving specific interactions between proteins, viral nucleic acid and specific membrane components, which is poorly understood.
We are particularly interested in biomembranes and membrane-related biological processes. For example, many toxins exert their damage on target cells by interfering with membrane functions. Misfolded peptides affect membranes of neuronal cells so that the communication between these cells is abolished; diseases such as Alzheimer's or Parkinson's are the consequence. As a further example, viral assembly, such as that of the Human Immunodeficiency Virus occurs at the plasma membrane of host cells and is a complex process involving specific interactions between proteins, viral nucleic acid and specific membrane components, which is poorly understood.
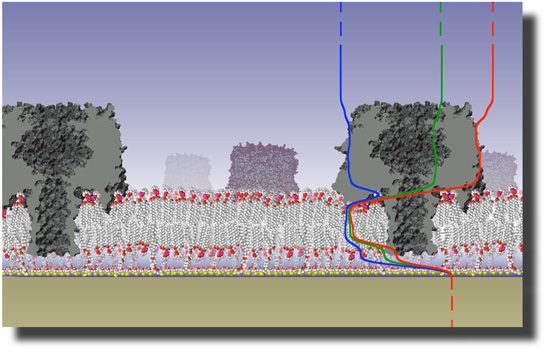
Traditional biological tools have been invaluable in identifying the molecular components of biomolecular assemblies and determining their qualitative functions. Yet, an integrated understanding of the underlying mechanisms and principles, as well as their dynamic function is largely incomplete.
Physics contributes to this understanding on many levels, from providing a theoretical framework for the quantitative description of Soft Condensed Matter to providing experimental characterization tools for probing the dynamics of function, assembly and disassembly.
Progress in the understanding of supramolecular complexes will impact science and technology areas far beyond biology. Already to date it exerts a significant impact on materials engineering and biotechnology, and is likely to extend into areas yet unforeseen.
News
October 2015
Research article published in Structure. Heinrich et al. demonstrate that PTEN, a biomedically importa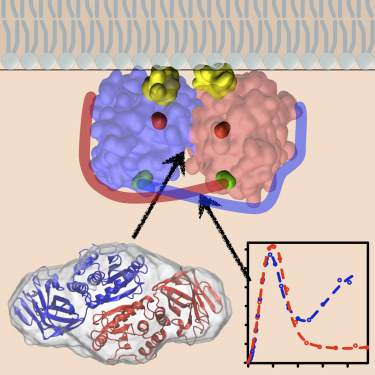 nt tumor suppressor, forms dimers in solution using small angle x-ray scattering. This finding may have important consequences for our understanding of PTEN function in the cell.
nt tumor suppressor, forms dimers in solution using small angle x-ray scattering. This finding may have important consequences for our understanding of PTEN function in the cell.
September 2014
Heinrich and Lösche publish ed a review article in BBA Biomembranes summarizing the significant progress over the last decade in structural biology using neutron reflectometry.
ed a review article in BBA Biomembranes summarizing the significant progress over the last decade in structural biology using neutron reflectometry.
May 2013
Collaborative Agreement between the NIST Center for Neutron Research and the Supramolecular Structures Lab renewed for a second term of five years. See more...
See more...
News Archive ...
Contact
SMSL,
c/o Amanda Bodnar
Physics Department
Wean Hall
5000 Forbes Ave
Pittsburgh PA 15213
USA
Tel. 412-268-8367
Fax 412-268-8252
We are a member of the joint UPSM-CMU MBSB graduate program.![]()