Biological Physics
Membrane Structure and Interactions Lab (MSIL)
Nagle and Tristram-Nagle Group
Research Highlights:
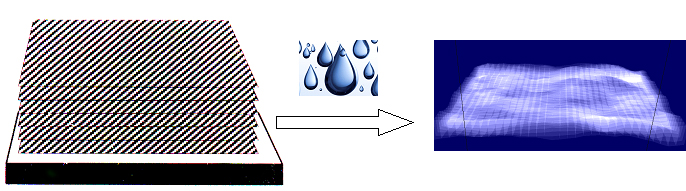
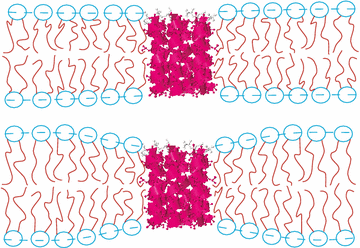
The Membrane Structure and Interactions Lab (also known as the Nagle and Tristram-Nagle Group) studies biomimetic membranes, using a variety of experimental techniques, including x-ray and neutron scattering, densimetry, calorimetry and some spectroscopy and light scattering, combined with physics theory and MD simulations (through collaborations). Cell membranes have an underlying structure called the lipid bilayer. We want to know why there are over 400 different lipids in red blood cell membranes, instead of just one, which is all that is needed to make a bilayer. Lipids are fairly small molecules with a split-personality, which we call amphiphilic. They get this name because they have one greasy end (hates water) and one polar end (loves water), which cause them to self-assemble into a bilayer arrangement where the greasy ends face each other and the polar “headgroups” face the water phase. Use of the hydrophobic effect was Nature’s solution to self-assembling small compartments (cells and organelles) which are containers for other important biomolecules, including DNA, RNA and proteins. Focus on Precise Structures: To the left is one of our experimental biomimetic samples, the unilamellar vesicle, On the left above is a cartoon of a stack of oriented bilayers, and the arrow indicates hydration through the vapor. You can see that a hydrated, individual bilayer is not perfectly flat; this is because when dry lipid bilayers take up as much water through a humid vapor as if they were under water, they fluctuate. (Getting stacks to fully hydrate through the vapor was at first difficult!) When a stack of bilayers fluctuates, it produces diffuse scattering when x-rayed. The Nagle lab was the first to be able to analyze the diffuse x-ray scattering data from fully hydrated, oriented, fluctuating bilayers. We had to modify the liquid crystal theory of Pierre-Gilles de Gennes and Alain Caillé. This took our lab a long time and it involved grad students Ruitian Zhang, Horia Petrache and Yufeng Liu, and postdoc Yulia Lyatskaya (see publications). From these data we obtain not only bilayer structure, such as precise lipid areas and thicknesses, but also the material properties, KC (bending) and B (compression) moduli. These moduli open a window to determining the very weak forces between membranes, such as van der Waals (attractive), hydration and fluctuation (repulsive) forces. A summary table of all of the lipid areas, thicknesses and bending moduli can be found here. We also find that each lipid is at equilibrium with a unique number of total water molecules. These values can be found in the citations in the summary table. We think it is important to study lipid membranes in their fully hydrated state because that is how they occur in the human body. In addition to studying the structure and properties of pure lipid bilayers, we have successfully added peptides (antimicrobial and HIV-related) and cholesterol to pure lipids. We have found that three peptides soften membranes, while cholesterol’s stiffening effect depends on the number of double bonds in the lipid chains. By collaborating with MD simulators and/or using a numerical fitting model of a bilayer electron density, we can determine the position of the peptide or cholesterol within the bilayer. The cartoon at the top left of this page shows the membrane thinning that occurs when a transmembrane peptide is signficiantly shorter than the underlying lipid membrane (lower image). See the research icons for more details about our research.
[The MSIL WEBsite was prepared with the Adobe Dreamweaver software.] |