Welcome to the Biophysics Group at CMU!

Biophysics is an exciting interdisciplinary frontier in physics. It applies tools and techniques of physics to understand how biological systems work, from the scales of single molecules to an entire living organism—and beyond. At the same time, biological phenomena offer us physicists unique opportunities to learn new physics on complex, non-equilibrium systems.
The Biophysics Group at Carnegie Mellon combines theory, experiments and computational modelling to investigate the fundamental principles that govern the structure, mechanics and dynamic behavior of living systems across different levels of biological organization and complexities.
For more details on our ongoing research activities, please have a look here.
Research Highlights
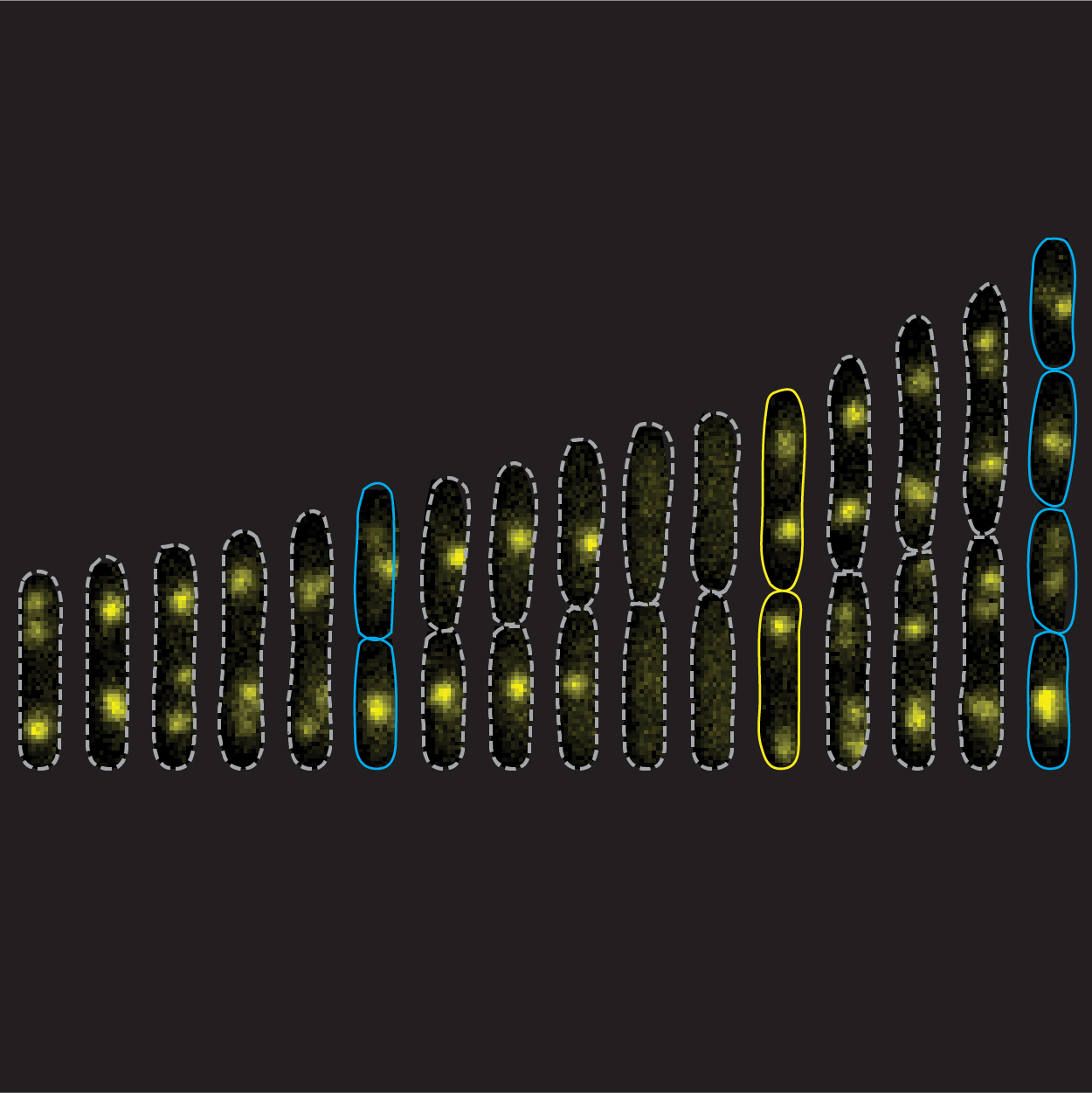

“Mechanistic origin of cell-size control and homeostasis in bacteria”, Fangwei Si, Guillaume Le Treut, John T. Sauls, Stephen Vadia, Petra Anne Levin, & Suckjoon Jun
Current Biology 29(11), 1760–1770.e7 (2019)
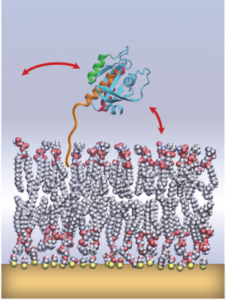
“Uncovering a membrane-distal conformation of KRAS available to recruit RAF to the plasma membrane.”, F. Heinrich, M. Lösche and collaborators
PNAS 117, 24258 (2020) (Commentary)
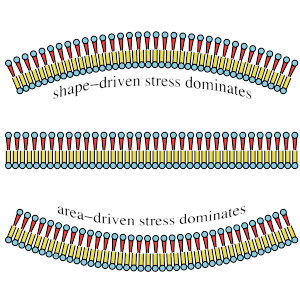 “Spontaneous curvature, differential stress, and bending modulus of asymmetric lipid membranes”, A. Hossein and M. Deserno.
“Spontaneous curvature, differential stress, and bending modulus of asymmetric lipid membranes”, A. Hossein and M. Deserno.
Biophys. J. 118, 624–642 (2020).
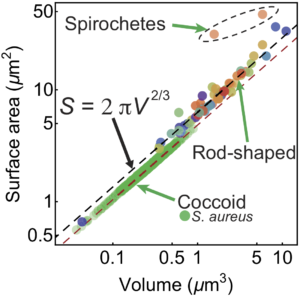 “Surface-to-volume scaling and aspect ratio preservation in rod-shaped bacteria”, N. Ojkic, D. Serbanescu and S. Banerjee.
“Surface-to-volume scaling and aspect ratio preservation in rod-shaped bacteria”, N. Ojkic, D. Serbanescu and S. Banerjee.
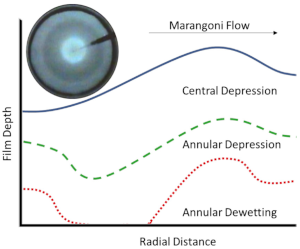 “Flow regime transitions and effects on solute transport in surfactant-driven Marangoni flows“, S.V. Iasella, N. Sun, X. Zhang, T.E. Corcoran, S. Garoff, T.M. Przybycien, R.D. Tilton.
“Flow regime transitions and effects on solute transport in surfactant-driven Marangoni flows“, S.V. Iasella, N. Sun, X. Zhang, T.E. Corcoran, S. Garoff, T.M. Przybycien, R.D. Tilton.
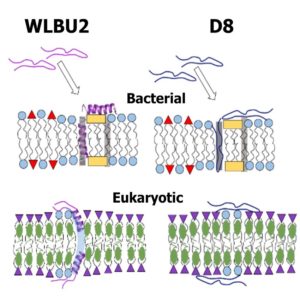 “Synergistic biophysical techniques reveal structural mechanisms of engineered cationic antimicrobial peptides in lipid model membranes”, F. Heinrich, A. Salyapongse, A. Kumagai, F.G. Dupuy, K. Shukla, A. Penk, D. Huster, R.K. Ernst, A. Pavlova, J.C. Gumbart, B. Deslouches,Y.P. Di, and S. Tristram-Nagle.
“Synergistic biophysical techniques reveal structural mechanisms of engineered cationic antimicrobial peptides in lipid model membranes”, F. Heinrich, A. Salyapongse, A. Kumagai, F.G. Dupuy, K. Shukla, A. Penk, D. Huster, R.K. Ernst, A. Pavlova, J.C. Gumbart, B. Deslouches,Y.P. Di, and S. Tristram-Nagle.

 Banerjee
Banerjee Deserno
Deserno Si
Si Heinrich
Heinrich Garoff
Garoff Nagle
Nagle Tristram-Nagle
Tristram-Nagle