Search:
Browse directory
Filter
- All
- air quality and climate
- biotechnology and pharmaceutical engineering
- catalysis and surface science
- energy, decarbonization, and sustainability
- process systems engineering
- soft materials and complex fluids
Faculty
-
-
-
-
-

Neil Donahue
Thomas Lord Professor
Director, Steinbrenner Institute for Environmental Education and Research -

Andrew Gellman
Lord Professor
Co-Director, Wilton E. Scott Institute for Energy Innovation
-
-
-

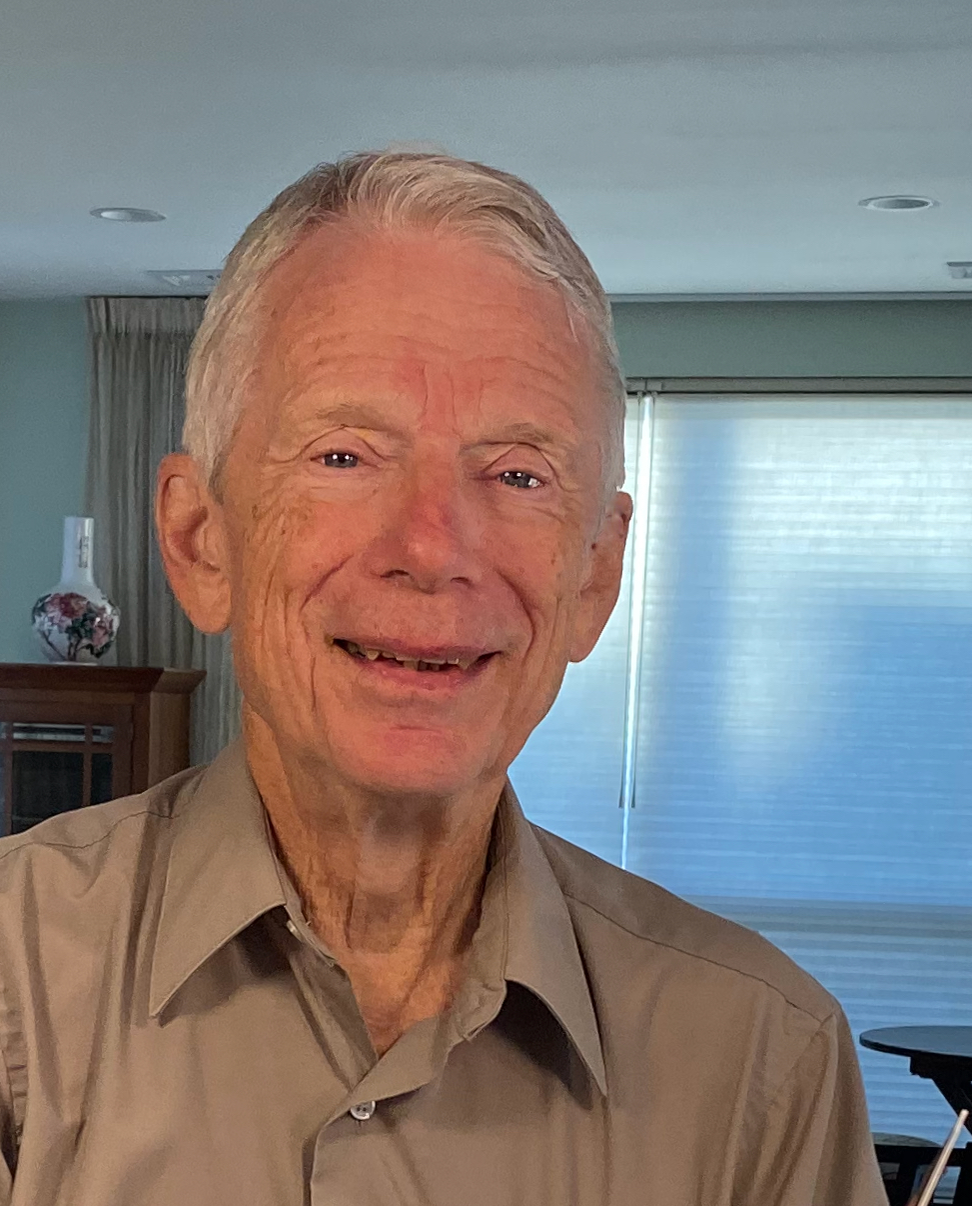
Ignacio Grossmann
Rudolph R. and Florence Dean University Professor
-
-
-
-
-
-
-
-
-
-
Courtesy & Adjunct Faculty
-
-
-

Debangsu Bhattacharyya
GE Plastics Material Engineering Professor, Department of Chemical and Biomedical Engineering, West Virginia University
Adjunct Faculty -
-
-

Krzysztof Matyjaszewski
J. C. Warner University Professor of Natural Sciences
-
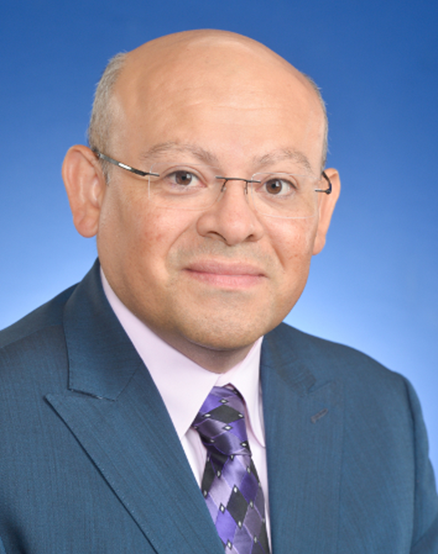

Salvador Garcia Muñoz
Senior Engineering Advisor, Eli Lilly and Company
Adjunct Faculty -
-
-
-
-
-
-
-
-
-
Emeritus
Staff
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-

Samantha Wessel
Manager of Administration and Assistant to the Department Head
-